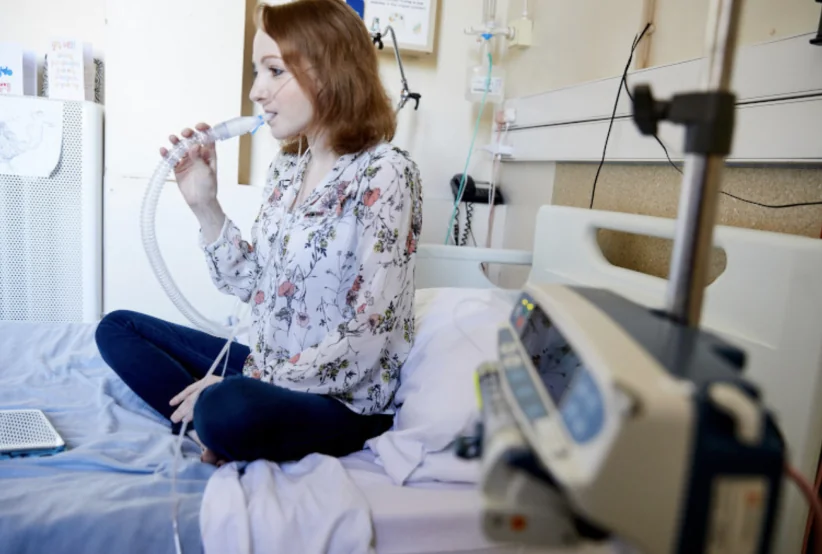
Biorepository Network
Launched in 2021, the CF AMR Syndicate biorepository network (formerly the UK CF Infection Biorepository) was established to advance translational science by improving access to high-quality, clinically relevant CF and non-CF samples, data, and expertise. By addressing a key challenge faced by researchers in industry and academia, the network is accelerating the development of new treatments for CF-related and other chronic lung infections.
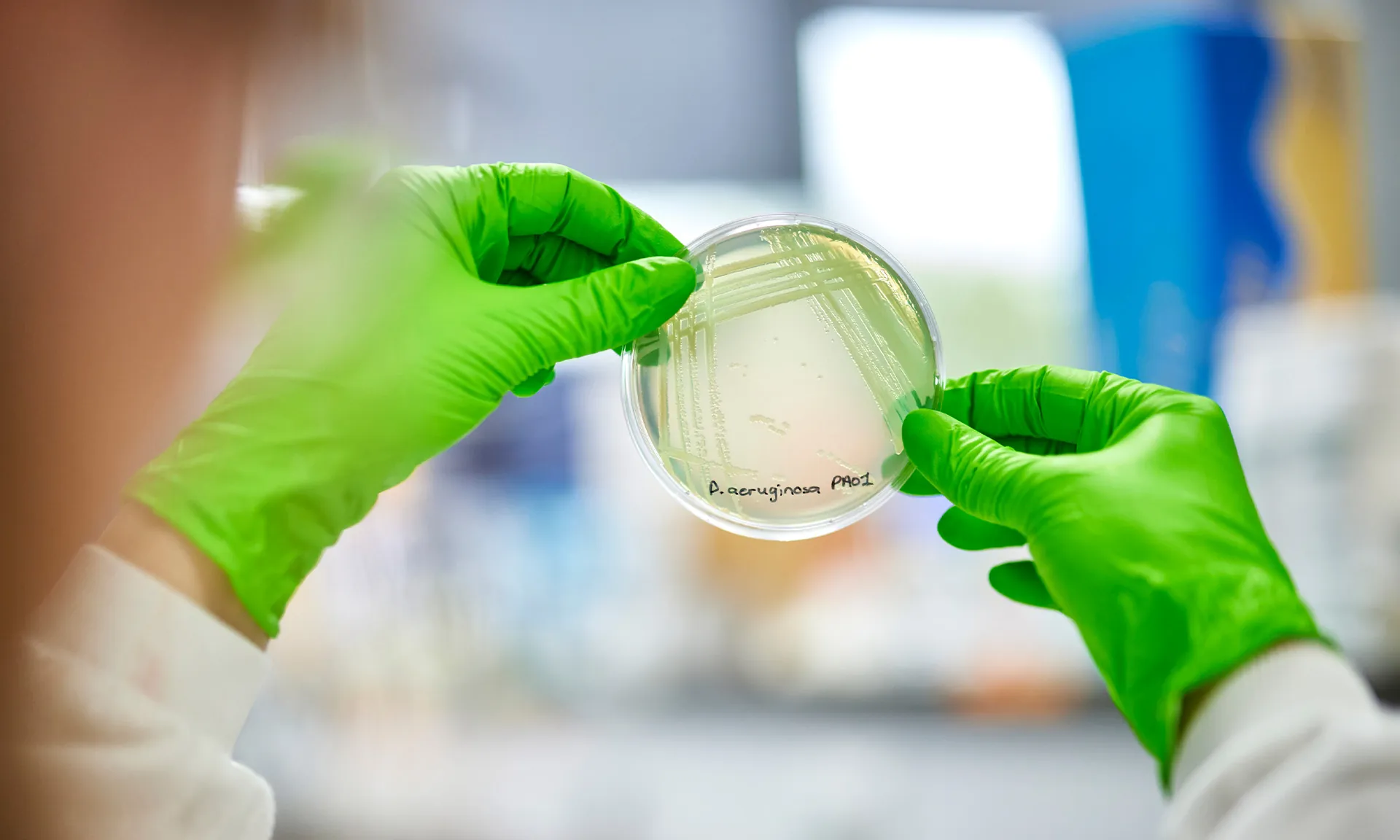
About the network
Funding from the Cystic Fibrosis Trust, Antabio, and the Cystic Fibrosis Foundation enabled the launch of the biorepository network in 2021. By making it easier for researchers to access high-quality clinical samples, associated data and expert guidance, this resource enables researchers to translate their discoveries into effective new treatments faster.
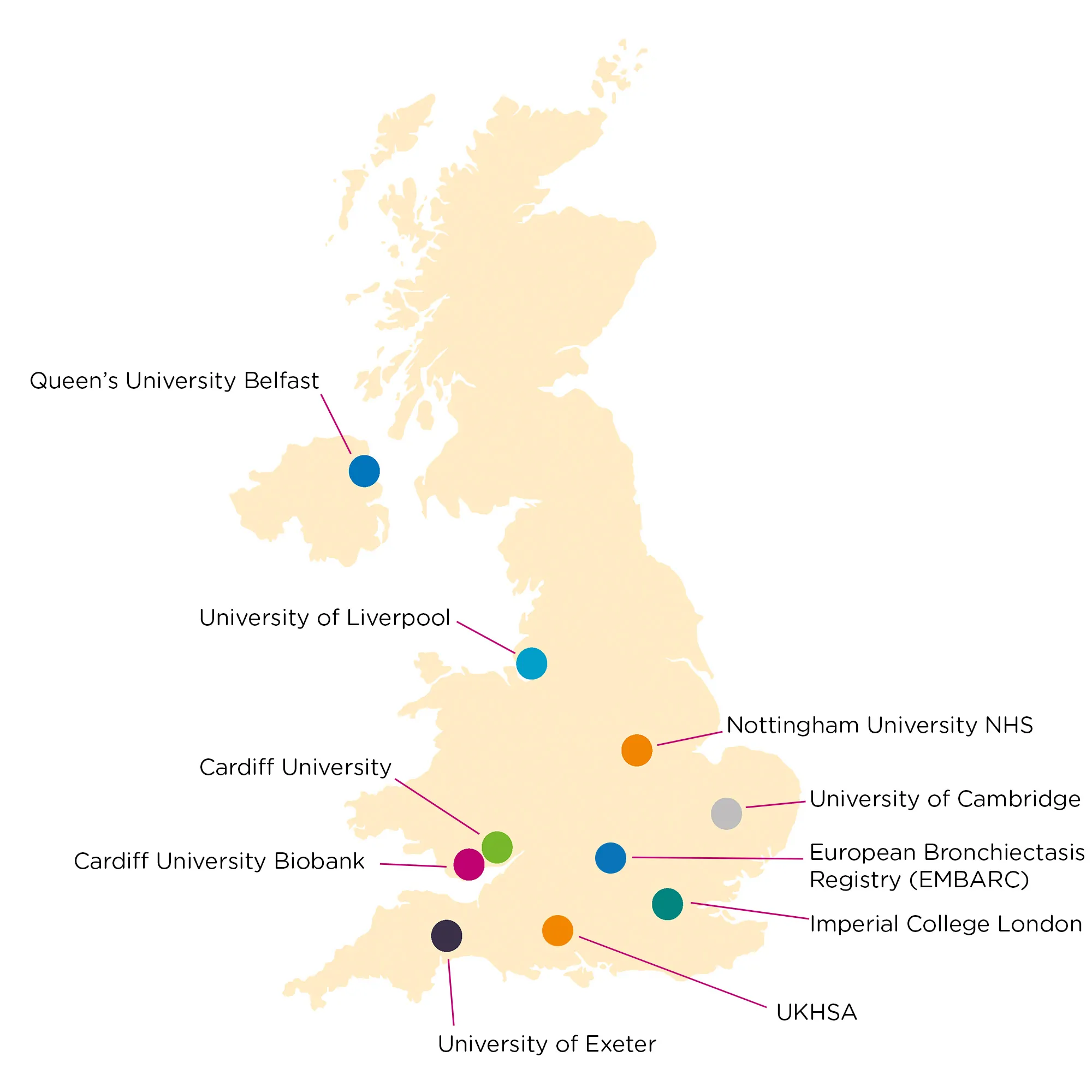
How the Network Works
- The biorepository runs as a coordinated national network of UK labs, each with existing collections and unique capabilities.
- A standard Material Transfer Agreement (MTA) is in place across several centres in the network, speeding up the time the time from enquiry to sample receipt.
For more information, read the laboratory descriptions here
The biorepository supplies samples and engages in collaborative projects with academic and industry researchers in the UK and globally.
Requests for samples, data, or capabilities can be sent directly to the relevant centre within the network, with terms dependent on the nature of the request. Requests may be subject to a fee depending on the nature and source of the request. See our FAQs for more information
If you’re unsure which centre to contact, need help contacting them, or have a broader enquiry about the biorepository or the CF AMR Syndicate, please get in touch with us with your query and we can connect or signpost you to the right people
Please also see FAQs
Resources Available to Researchers
a) Non-CF and CF-reference strains and strain panels for common pathogens of the CF lung including Pseudomonas aeruginosa, Burkholderia cepacia complexand non tuberculous mycobacteria (NTM).
b) Bacterial pathogens including Strenotrophomonas maltophilia, Achromobacter species, Staphylococcus aureus, Streptococcus spp, Haemophilus Influenzae,
c) Strains and strain panels are well-characterised, with linked phenotypic data, whole-genome sequences, and associated published research.
d) Fungal and other Aspergillus species, Exophiala, Rasomsonia. Many of the clinical isolates are genome sequenced with clinical metadata associated.
e) Contemporary post-modulator isolates reflecting current pathogen populations are also available.
f) Isolates are also available from both longitudinal studies and as multiple isolates from individual patients.
a) Frozen or whole samples with linked clinical metadata.
b) Signposting to centres to support prospective collection of sputum
3. Antimicrobial susceptibility testing, including combination testing in different formats and models.

Designed in collaboration with experts from the biorepository network, this diverse mini-panel combines clinically derived isolates and reference strains to generate robust, clinically relevant preclinical data that reflects real-world pathogen diversity and resistance patterns, helping innovators efficiently assess potential therapeutics..
The panel includes strains of P. aeruginosa, M. abscessus, and B. cepacia, carefully selected to represent the genomic diversity of CF pathogens.
The mini panel is now available to support studies through the UK Health Security Agency’s (UKHSA) Open Innovation in AMR platform.
Developed by experts from the biorepository network, this guidance document provides best-practice recommendations for selecting strains in efficacy testing. Researchers working on early-stage therapeutics or looking to develop a CF data package will find this paper valuable because it:
- Provides guidance for selecting clinically relevant CF pathogen strains, including P. aeruginosa and M. abscessus, to design robust preclinical efficacy panels that reflect genetic and phenotypic diversity
- Highlights gaps in traditional lab strains demonstrating why disease relevant isolates are essential for meaningful preclinical results.
- Summarises available strain collections and panels, to help identify resources to support testing and therapeutic development.
- Supports reproducible research, emphasising strain selection that allows validation and comparability across studies.
Access the paper here
Read the report here.
Meet the network
Cardiff University Biobank (CUB) is a dedicated human tissue storage facility which aims to provide high quality biosamples to patient benefitting research. CUB provides ethical and logistical provision for the collection of samples from Cystic Fibrosis and Respiratory patient cohorts within Cardiff & Vale University Health Board. We work closely with the clinical stakeholders of these collections to support their research interests, while facilitating sample access to the wider academic and commercial research landscape of the UK and abroad.
Collections/Samples available
Plasma, Serum, Whole blood, Sputum, Urine, Cough swabs
Data:
Basic clinical and demographic
Main contact:
Edward Gait-Carr, Cardiff University
CUBiobank@cardiff.ac.uk
EMBARC is a multinational biorepository for bronchiectasis collection sputum, blood, urine and associated bacterial isolates. The collection includes >20,000 organisms, with a predominance of Pseudomonas aeruginosa. The collection has permissions for sharing including with commercial organisms and isolates have associated clinical meta-data which can be shared with appropriate data sharing agreements in place. The vast majority are from patients without cystic fibrosis.
EMBARC seeks collaborations with partners to advance science in the field of bronchiectasis.
Collections/Samples available:
- EMBARC biobank collection including aeruginosa, H.influenza plus some other 50 organisms
Data
Comprehensive and whole genome sequence
Main contact and details:
James Chalmers, University of Dundee
Queen’s University Belfast has an extensive collection of over 10,000 clinical respiratory isolates from sputum, BALF, cough swabs, and saliva. Isolates include key pathogens such as Pseudomonas aeruginosa, Staphylococcus aureus and anaerobes such as Prevotella. Isolates are from longitudinal and cross-sectional studies in CF, bronchiectasis, COPD, and healthy controls, with associated phenotypic and genomic data.
The centre offers expertise in quantitative sputum microbiology, anaerobic culture, MALDI-TOF, qPCR, and 16S rRNA sequencing. It serves as a central lab for clinical studies, with robust sample tracking and antimicrobial testing using standard assays and under conditions that replicate the lung environment.
Collections/samples available
- US-Ireland Study, Longitudinal and Individual, 2010-2013 PMID: 30905581, 29946004 CF:
- Stable and exacerbation study, 2005-2008 PMID: 18263800, 26774156
- IMI Collection PMID: 33367642, 31767727, 36006824, 35730434, 35549675, 34463860, 31762328
- iBEST collection PMID: 32855225.
- BronchCF (QUB & University College Cork) with Isolates from participants pre- and post-modulator
- Samples available across the above collections include aeruginosa, B.cepacia complex, Achromobacter spp. H.influenzae S.aureus, S. maltophilia Streptococcus spp, Prevotella spp, Ralstonia spp. and several other pathogens.
Data
Some comprehensive, whole genome sequencing, sensitivity and clinical metadata across the collections
Other capability available:
Queen’s University Belfast has expertise in conventional quantitative sputum microbiology, culture of anaerobic bacteria and molecular detection and quantification of pathogens using MALDI, quantitative PCR assays and next-generation sequencing. The centre acts as a central laboratory for quantitative sputum microbiology in clinical studies with sample tracking and specimen management systems in place.
The centre also has extensive expertise in antimicrobial susceptibility testing including evaluation of the efficacy of novel antimicrobial compounds alone and in combination. A range of assays can be used including testing under conditions which mimic in vivo conditions in the lung such as growth in artificial sputum media, biofilm and under low oxygen or anaerobic conditions.
Main contact: Michael Tunney, Queen’s University Belfast
Our Open Innovation program offers researchers and innovators unprecedented access to world-class expertise, facilities, and collaborative opportunities.
Cutting-edge screening: Utilise our state-of-the-art screening cascade to evaluate a wide range of novel antimicrobial therapies, including non-traditionals.
Expert training and support: Gain hands-on experience and technical expertise through training and direct access to our renowned microbiologists at Porton Down laboratories.
Strategic partnerships: Leverage our existing network of interdisciplinary partners to access support for innovative antimicrobial research projects.
Samples/ collections available
- CF AMR Syndicate testing panel covering P. aeruginosa; M.abscessus and B. cepacia. This panel is comprised of a set of well characterised strains and CF-isolates relevant to the CF lung and representative of the genomic diversity of the pathogens of interest
- Panels of contemporary MDR K. pneumoniae, E. coli, A. baumannii and P. aeruginosa co-designed with the PACE consortium (Medicines Discovery Catapult, LifeArc, Innovate UK)
- Wider panels of ESKAPE pathogens, candidal species and other relevant human pathogens including N. gonnorrhoeae and NTMs.
Data
All strains are fully whole genome sequenced, with associated antibiogram, sequence types and in most cases the indication source, geographic location and year of isolation.
Main contact
Charlotte Hind, UKHSA
The Floto group research is focused on the mechanisms of immune cell interaction with bacteria, and how intracellular killing and inflammation are regulated and modified during bacterial lung infection. We are investigating how bacteria evolve and adapt to improve their survival by functional characterisation of the key drivers of pathoadaptation, including integration of genomic, transcriptomic and phenotypic data employing GWAS, multi-omics integration and causal learning methods. We are also engaged in antibiotic discovery using structure-guided approaches and metabolic stability of drug-like molecules. We are also developing home based monitoring of disease stability and exacerbation predictions in cystic fibrosis and bronchiectasis patients.
Collections available:
- Cambridge NTM collection
- Cambridge aeruginosacollection
Data: Comprehensive and whole genome sequencing data available
Main contact: Andres Floto, University of Cambridge
The Nottingham University Hospitals Pathogen Bank is a curated collection of clinically relevant bacterial and yeast isolates. Developed to aid NHS laboratories, academia and industry partners to validate new and existing diagnostic assays, drug development and for use in research. There are currently approximately 4900 bacterial and yeast strains comprising of over 130 genera. This collection includes 313 isolates collected from sputum and cough swabs from people with CF.
We have expertise in clinical diagnostics including antimicrobial identification and sensitivity testing. Options for strain only partnerships or collaborations are available.
Collections/samples available
- Nottingham bacterial isolate collection incl: aeruginosa Achromobacter spp, Burholderia spp, H.influenzae, E.coli, Moraxella, Serratia, S.maltophilia
- Nottingham fungal isolate collection incl: Aspergillus, Candida, Exophiala, Scedosporium
Data
Antimicrobial susceptibility data
Other capability:
Microbiological culture – aerobic, anaerobic, containment level 3 facilities.
Antibiotic sensitivity testing – broth dilution, disc diffusion, E-test.
Bacterial identification – MALDI-TOF, 16S Sanger Sequencing
Main contact
Michelle Lister, Nottingham University Hospitals NHS Trust
Email: nuhnt.pathogenbank@nhs.net
The National Heart and Lung Institute (NHLI) at Imperial College London has been collecting samples from people with CF from the Royal Brompton and Harefield hospital in Chelsea, London since 2014. Initially curating P.aeruginosa isolates only, the collection has since expanded to collecting a variety of CF-relevant pathogens, including fungal species such as Candida spp. We house longitudinal samples, with samples collected from single patients spanning up to ten years – these samples are vital for studying bacterial evolution and chronicity.
Imperial joined the Biorepository network upon its creation in 2021, bringing expertise in microbiology and bacteriology. We encourage collaborations with clinical/ fundamental science researchers, biotech and pharma.
Collections/samples available
- London fungal collection incl Apergillus, Exophiala and Rasomsonia
- London CF Bacterial Isolate Collection incl Achromobacter , Bcepacia, P.aeruginosa (mucoid and non mucoid) S. aureus, Stenotrophomonas Maltophilia, Klebsiella pneumoniae, MRSA
- London non CF bronchiectasis isolate collection aeruginosa
Data
Comprehensive, clinical, antimicrobial susceptibility testing & whole genome sequencing data
Other capability:
Collaborations for testing of novel antimicrobial compounds against CF-relevant pathogens
Main Contact
Amie Micallef, National Heart and Lung Institute, Imperial College, London.
FAQs
General
The biorepository network (previously known as the UK CF Infection Biorepository) was launched in September 2021 to address a key challenge in CF antimicrobial discovery and development: the difficulty researchers face in accessing all the sample types they need for preclinical testing.
The biorepository runs as a coordinated national network of UK labs, each with existing biobank collections and capability. Laboratories in the network are distributed across the University of Belfast, University of Cambridge, University of Cardiff, University of Exeter, University of Liverpool, Imperial College London, Nottingham University Hospitals NHS Trust, and the University of Dundee.
The biorepository supplies samples and runs collaborative projects with academic and industry researchers in the UK and globally.
Funding from the Cystic Fibrosis Trust, Antabio, and the Cystic Fibrosis Foundation enabled the establishment of this resource.
Available Samples, Data and Capability
- Non-CF and CF-reference strains and strain panels for common pathogens of the CF lung including Pseudomonas aeruginosa, Burkholderia cepacia complex and non tuberculous mycobacteria (NTM). Strain panels are associated with published data including genome sequence;
- Bacterial and fungal strains isolated from the lungs of people with CF in the UK; aeruginosa, Achromobacter xylosoxidans, M.abscessus, B. Cepacia, Stenotrophomonas maltophilia, S. aureus, Streptococcus spp, Aspergillusspecies, Exophila, Rasomsonia. Many are genome sequenced with associated patient metadata and include recent isolates, collections from longitudinal studies and multiple isolates from single sputum samples.
- Sputum (frozen, whole) with associated patient metadata;
- Signposting to centres to support prospective collection of sputum;
- Antimicrobial susceptibility testing including combination testing in different formats and models.
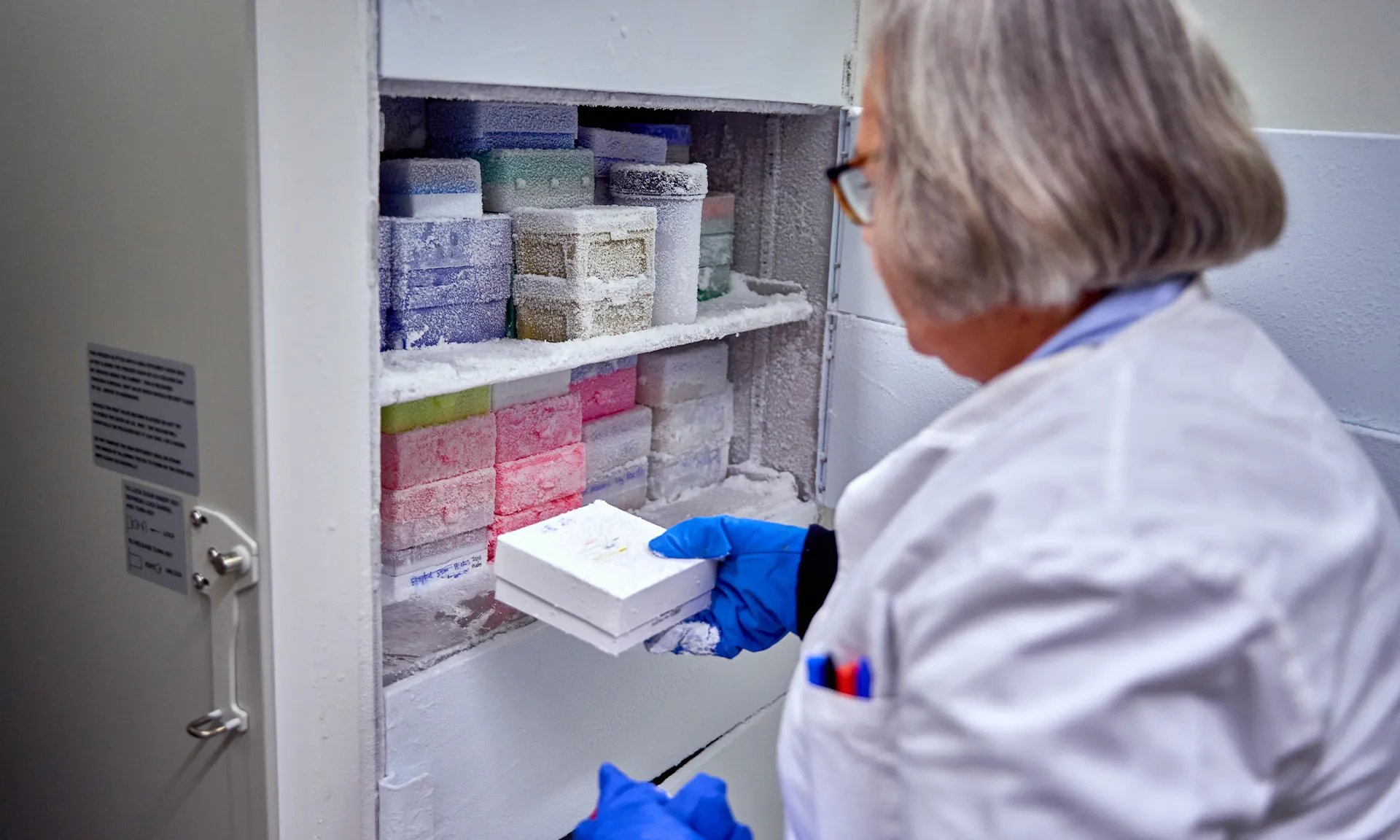
Pure isolates are typically stored long term in tryptic soya broth with 8% DMSO or using microbeads both at -80ºC.
Frozen sputum is cut and stored frozen at -80ºC, some spun supernatant also stored at -80 ºC is also available (most utility for investigation of inflammatory markers).
BAL spun supernatant is stored at -80ºC (most utility for investigation of inflammatory markers).
Each biobank and collection centre in the network has in place their own ethical consent governed by local biobank ethical review committees.
This depends on the sample but can include; patient characteristics, Spirometry (FEV1), medication including long-term antibiotics, sample type, clinical state and microbiological status.
In many cases, yes.
Yes, a number of centres in the network have expertise in antimicrobial susceptibility testing, including combination testing, in different formats and models. Depending on the centre, this might be available on a fee-for-service or collaborative basis.
Yes, please see this published strain guidance document for more information.
The Cystic Fibrosis Foundation has links to a number of antimicrobial tools and resources including clinical isolates from people with CF living in the US.
The Canadian Burkholderia cepacian Complex Research and Referral Laboratory also holds a number of isolates from people with CF living in Canada.
Strains from published reference panels are available from the Belgium Coordinated Collection of Microorganisms/Laboratory for Microbiology
For general antibiotic resistance isolates the AR Isolate Bank and the Public Health England National Collection of Type Cultures NCTC , the National Collection of Pathogenic Fungi NCPF and the National Collection of Pathogenic Viruses NCPV The human microbiome project repository also makes organisms and DNA available to the community.
Processes
Requests for samples, data, or capabilities can be sent directly to the relevant centre within the network, with terms dependent on the nature of the request. Requests may be subject to a fee depending on the nature and source of the request. See our FAQs for more information
If you’re unsure which centre to contact, need or help contacting them, or have a broader enquiry about the biorepository network or the CF AMR Syndicate, please get in touch with us with your query and we can connect or signpost you to the right people
Yes, please make an enquiry through the website and a member of the team will discuss with you.
If you are interested in donating your samples for research, please discuss this with your CF centre. Unfortunately, the biorepository does not accept individual donations.
To help cover a proportion of the administrative running costs of centres, cost recovery fees are in place and payments will be managed directly by each of the centres.